How DNA is Sequenced
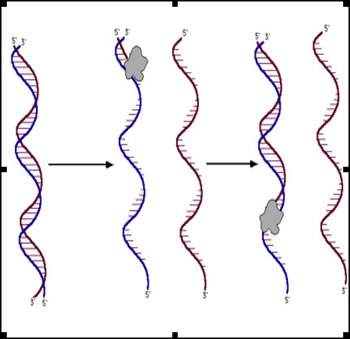 |
DNA sequencing reactions are just like the PCR reactions for replicating DNA. The reaction mix includes the template DNA, free nucleotides, an enzyme (usually a variant of Taq polymerase) and a 'primer' ---- a small piece of single-stranded DNA about 20-30 nt long that can hybridize to one strand of the template DNA. The reaction is initiated by heating until the two strands of DNA separate. Then the primer anneals/binds to its intended location and DNA polymerase starts elongating the primer. If allowed to go to completion, a new strand of DNA would be the result. If we start with a billion identical pieces of template DNA, we'll get a billion new copies of one of its strands. |
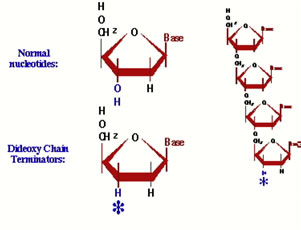 |
Dideoxynucleotides: We run the reactions, however, in the presence of a dideoxyribonucleotide. This is just like regular DNA, except it has no 3' hydroxyl group --- once it is added to the end of a DNA strand, there is no way to continue elongating it. Now the key to this is that MOST of the nucleotides are regular ones, and just a fraction of them are dideoxy nucleotides.... |
 |
Replicating a DNA strand in the presence of dideoxy-T: MOST of the time when a 'T' is required to make the new strand, the enzyme will get a regular one and there is no problem. MOST of the time after adding a T, the enzyme will go ahead and add more nucleotides. However, 5% of the time, the enzyme will get a dideoxy-T, and that strand can never again be elongated. It eventually breaks away from the enzyme, a dead-end product. Eventually, ALL of the copies will be terminated by a T, but each time the enzyme makes a new strand, the place it stops will be random. In millions of starts, there will be strands stopping at every possible T along the way. |
ALL of the strands we make started at one exact position. ALL of them end with a T. There are billions of them ... many millions at each possible T position. To find out where all the T's are in our newly synthesized strand, all we have to do is find out the sizes of all the terminated products! |
|
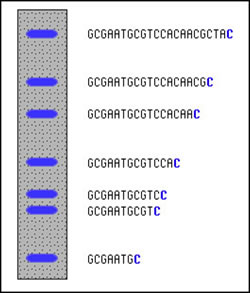 |
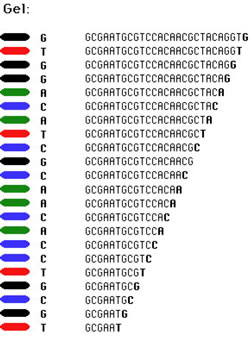
Determining fragment sizes: Gel electrophoresis can be used to separate the fragments by size. In the cartoon at left, we depict the results of a sequencing reaction run in the presence of dideoxy-Cytidine (ddC). First, the dideoxy nucleotides in my lab have been chemically modified to fluoresce under UV light. The dideoxy-C, for example, glows blue. With the reaction products on an 'electrophoresis gel', you will see something similar to that depicted at left. Smaller fragments are at the bottom, larger at the top. The positions and spacing shows the relative sizes. At the bottom is the smallest fragment that has been terminated by ddC; that is probably the C closest to the end of the primer (which is omitted from the sequence shown). |
Simply by scanning up the gel, we can see that we skip two, and then there is two more C's in a row. Skip another, and there is yet another C. And so on, all the way up. We can see where all the C's are in the sequence. |
|
 |
Putting all four deoxynucleotides into the picture: The spacing between the bands is not all that easy to figure out if just looking at a single nucleotide (e.g., C). Imagine, though, that we ran the reaction with *all four* of the dideoxy nucleotides (A, G, C and T) present, and with *different* fluorescent colors on each. NOW look at the gel we obtain (at left). The sequence of the DNA is rather obvious if you know the color codes ... just read the colors from bottom to top: TGCGTCCA-(etc...). (NOTE: Black is used here because it shows up better than yellow). |
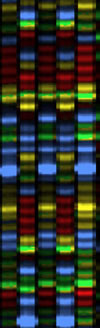 |
An Automated sequencing gel: That is exactly what we do to sequence DNA ---- we run DNA replication reactions in a tube, but in the presence of trace amounts of all four of the dideoxy terminator nucleotides. Electrophoresis is used to separate the resulting fragments by size and we can 'read' the sequence from the gel, as the colors march past in order. In a large-scale sequencing lab, we use a machine to run the electrophoresis step and monitor the different colors as they pass across a laser. Since about 2001, these machines --- not surprisingly called automated DNA sequencers --- have used 'capillary electrophoresis', where the fragments are piped through a tiny glass-fiber capillary during the electrophoresis step, and they come out the far end in size-order. There is an ultraviolet laser built into the machine that shoots through the liquid emerging from the end of the capillaries, checking for pulses of fluorescent colors to emerge. There might be as many as 96 samples moving through as many capillaries ('lanes') in the most common type of sequencer. |
Above is a screen shot of a real fragment of sequencing gel (this one from an older model of sequencer, but the concepts are identical). The four colors red, green, blue and yellow each represent one of the four nucleotides. The actual gel image, if you could get a monitor large enough to see it all at this magnification, would be perhaps 3 or 4 meters long and 30 or 40 cm wide. |
|
A 'Scan' of one gel lane: We do not even have to 'read' the sequence from the gel - the computer does that for us! Below is an example of what the sequencer's computer provides for one sample. This is a plot of the colors detected in one 'lane' of a gel (one sample), scanned from smallest fragments to largest. The computer even interprets the colors by printing the nucleotide sequence across the top of the plot. This is just a fragment of the entire sequence, which would span around 900 or so nucleotides of accurate sequence.
The sequencer also provides the operator a text file containing the nucleotide sequence, without the color traces.
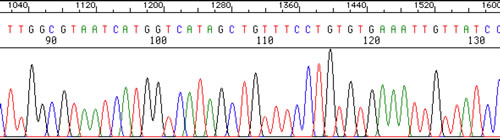
We can obtain the sequence of a fragment of DNA as long as 900-1,200 nucleotides using the above technology.